This collection of free browser-based tools supports common small tasks in genomics, PGT and sequencing data analysis.
All tools run entirely in your browser — files are never uploaded to our server.
They are intended for research use only and work with standard bioinformatics file formats.
Get in touch for further bioinformatics consulting or developement services.
Click any screenshot to enlarge it. Click a tool name in the menu or the Open button to launch it.
Look up the ISCN cytogenetic band for any genomic coordinate on the human or mouse reference genomes (hg19, hg38, mm39). Enter a chromosome and position to retrieve the cytogenetic band name or the other way around. Useful e.g. for annotating variants or CNV breakpoints with their cytogenetic location.
Open tool ↗
Count and visualise the number of sequencing reads aligned to each chromosome directly from a BAM file. Handles files up to 150 MB. Results are shown as a bar chart and a per-chromosome table. Useful for a quick sanity check of mapping distributions or for detecting gross mapping anomalies.
Open tool ↗
A collection of utilities for working with DNA sequences: reverse complement, GC content, melting temperature estimation, and more. Accepts free-text sequence input and returns results instantly in the browser.
Open tool ↗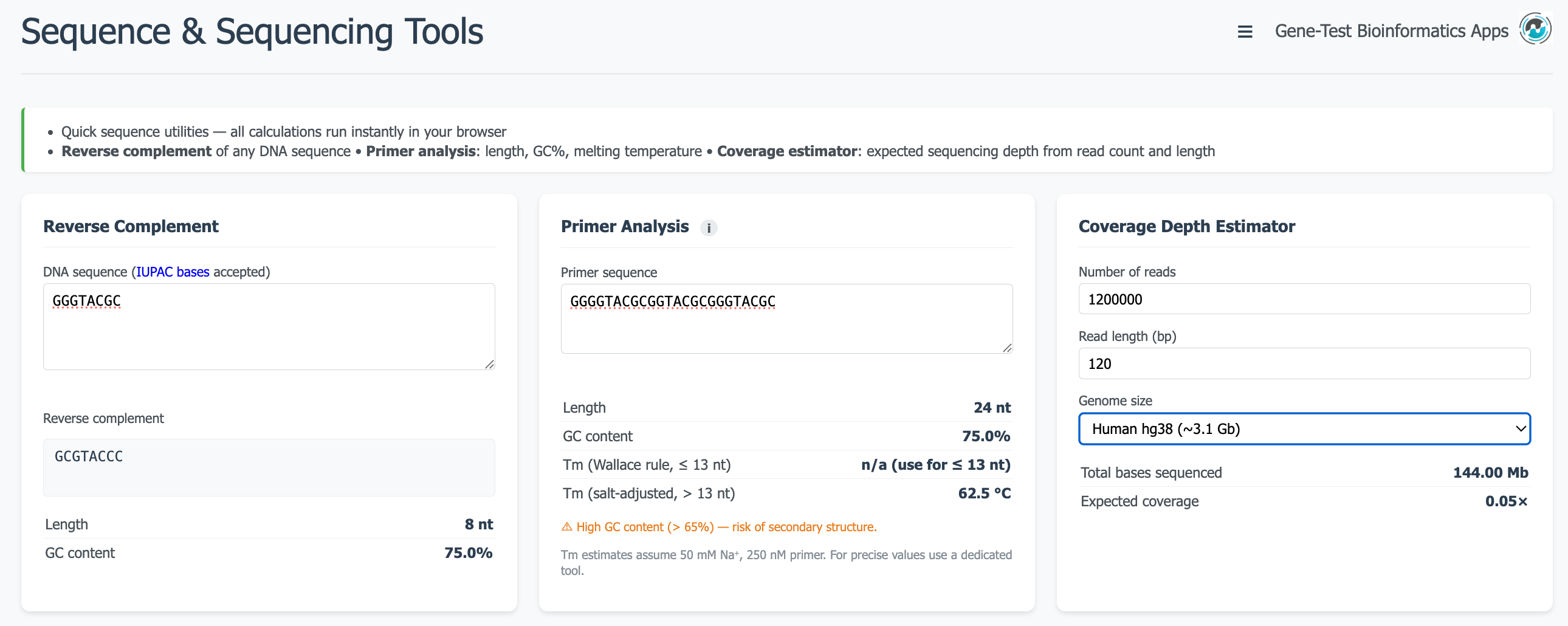
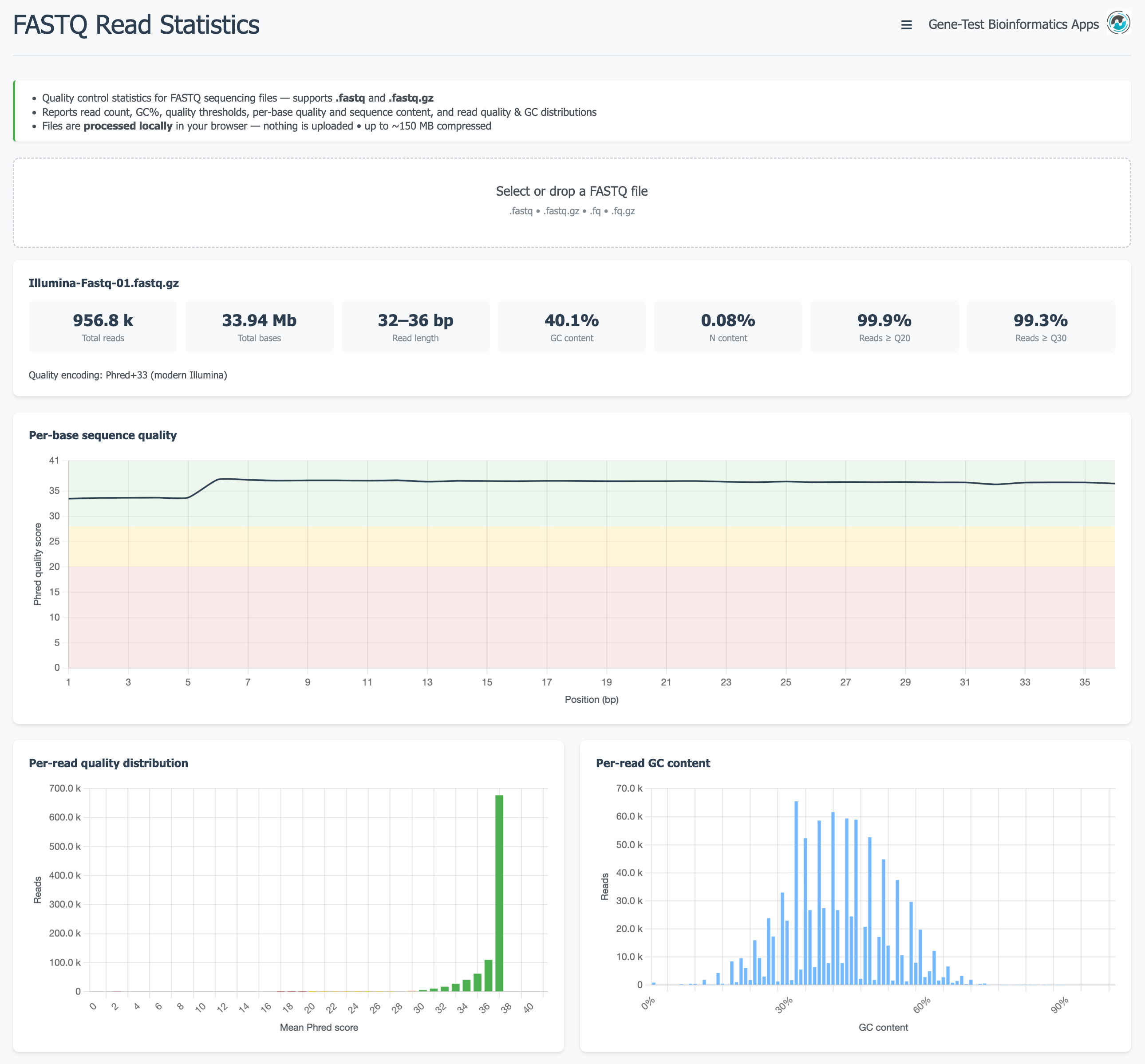
Comprehensive quality statistics for FASTQ sequencing files, including compressed
.gz files. Reports read count, length distribution, mean quality scores,
per-base nucleotide content, and GC distribution — similar in scope to FastQC,
running entirely in the browser without any installation.

Summary statistics for FASTA files: sequence count, total and per-sequence length,
N50/L50, GC content, ambiguous base fraction, and per-base nucleotide composition.
Supports standard .fasta / .fa / .fna files as
well as .gz-compressed variants. Suitable for assemblies, reference files,
or any multi-sequence FASTA.

Generate a chromosome idiogram with copy-number overlays for all 24 human chromosomes.
Two input modes: a karyotype string (e.g. +7 -13 +X(s)) for whole-chromosome
gains and losses, and a coordinate string for segment-level CNV calls.
Renders publication-style Giemsa-banded chromosomes using hg38 cytoband data.

